
Visualizing Multiclass Classification Results
Introduction
Visualizing the results of a binary classifier is already a challenge, but having more than two classes aggravates the matter considerably.
Let’s say we have $k$ classes. Then for each observation, there is one correct prediction and $k-1$ possible incorrect prediction. Instead of a $2 \times 2$ confusion matrix, we have a $k^2$ possibilities. Instead of having two kinds of error, false positives and false negatives, we have $k(k-1)$ kinds of errors. And not all errors are created equal: just as we choose an optimal balance of false positives and false negatives depending on the cost associated to each, certain kinds of errors in a multiclass problem will be more or less acceptable. For example, mistaking a lion for a tiger may be acceptable, but mistaking a tiger for a bunny may be fatal.
The goal of visualizing multiclass classification results is to allow the user to quickly and accurately see which errors are occurring and to start developing theories about why those errors are occurring; usually this would be to assist the user during iterative model development, but could also be used, for example, to communicate the behavior of a final classifier to a non-specialized audience.
In this article I employ two basic strategies to try and meet these goals: data visualization techniques and algorithmic techniques. It’s worth a quick reminder about why data visualization is valuable at all. The human visual system is extremely good at picking up certain kinds of patterns (generally those that correspond to spatial relationships and color), but is completely unable to see other kinds of patterns (the digits of $\pi$ coded as greyscale pixel brightness would look like pure noise) and worse yet has a tendency to see patterns in clouds of purely random data where none exist. A good visualization, then, ensures that any interesting structure in the underlying data will be presented in a way that is amenable to interpretation by the human visual system, while any irrelevant or statistically insignificant variation is suppressed.
Algorithmic techniques, on the other hand, do not rely on the human visual system’s ability to detect patterns, but automate the analysis that a human would have done anyway in some procedural way. Rather than merely making it easy to see where a relationship exists, an algorithmic solution would explicitly enumerate and rank the kinds of things the user is interested in. This approach can scale to large data sets much more efficiently, but requires us to trust the algorithm. Both data visualization and algorithmic techniques are useful in practice and are often best when combined.
The underlying problem is very open ended and I do not claim to have come up with any definitive solution, but I did find several novel and useful techniques that seem to me to be worth sharing.
Case Study
To explore the problem, we need some data and a toy classifier to work with.
The first step is to train and fit some reasonably good but not perfect classifier to some dataset that is reasonably amenable to classification but not linearly separable.
To meet these requirements, I chose the MNIST handwritten digit dataset; This data set is popular for testing multiclass classification algorithms and has the advantage of having very intuitive classes. The MNIST problem is to classify 28x28 gray scale images, which represent center and scaled images of handwritten digits, and assign them to one of ten classes, namely the digits 0 through 9. The MNIST data come pre-labeled and therefore ready to be fed into a supervised learning algorithm. Best of all, it is extremely easy to obtain in an R-friendly format due to its popularity.
As a preprocessing step, we will use T-SNE algorithm provided by the
tsne package to reduce the 784 dimensions of the raw pixel data to just
two dimensions, which we will simply call x and y.
mnist_tsne <- tsne(as.matrix(mnist_r10000[,1:784]))
xy <- as.data.frame(mnist_tsne)
colnames(xy) <- c('x', 'y')
xy$label <- mnist_r10000$Label
Next, we will apply a multinomial classifier from the nnet package (despite the name, the package actually provides MLP and multinomial log-linear models)
model <- multinom(
label ~ I(x^3) + I(y^3) + I((x^2)*y) + I(x*y^2) + I(x^2) + I(y^2) + x * y,
data=xy,
maxit=500
)
xy$prediction <- predict(model)
hits_and_misses = xy[xy$label != xy$prediction | rep_len(c(TRUE,FALSE), nrow(xy)),]
This model does an OK but not stellar job of classifying digits, achieving an overall accuracy of about 95%. This is what we want - a higher quality model would have too few misses to analyze deeply, while a simpler model wouldn’t be realistic enough to make a good case study.
Off-the-Shelf Pairs Plot
A good place to start with any dataset is a so-called “pairs” plot: a grid of plots showing relationships between every every possible pair of columns. The GGally package provides a particularly high-quality pair plot, so let’s start with that.
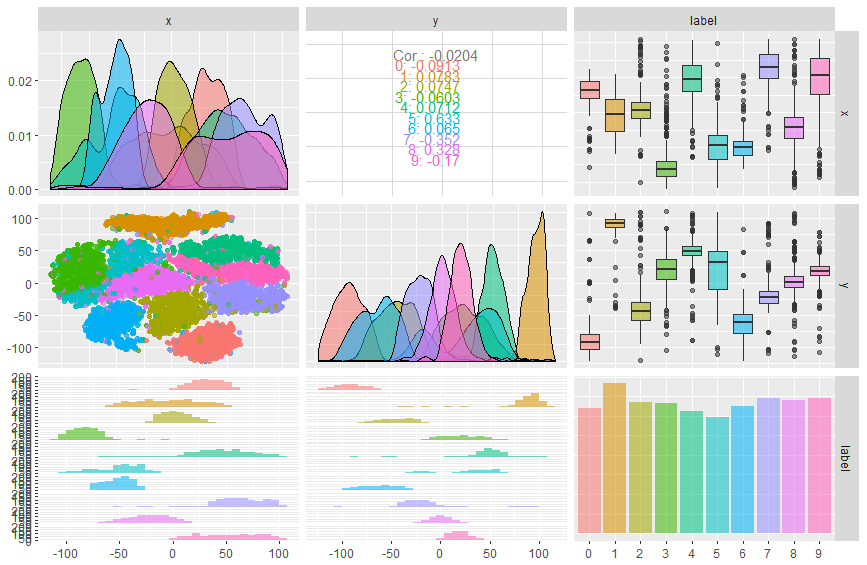
This plot was easy to create, but most of the relationships turn out to be
uninteresting, except in a negative sense: we can tell from the large amount of
overlap between classes in the univariate kernel density plots that neither x
nor y alone is able to classify digits very well. However, the leftmost plot
in the middle column, which shows a scatter plot of x and y color-coded
with the true class label, suggests that using both dimensions together with a
non-linear classifier may be effective.
Scatter Plot
To explore that further, let’s create a full-size scatter plot on x and y.
To compactly and intuitively represent both the true class and the
predicted class in the same plot, we will plot each point with the glyph
representing the true class and a color representing the predicted class. To
avoid overwhelming the plot, we plot only a random sample of 1,000 points.
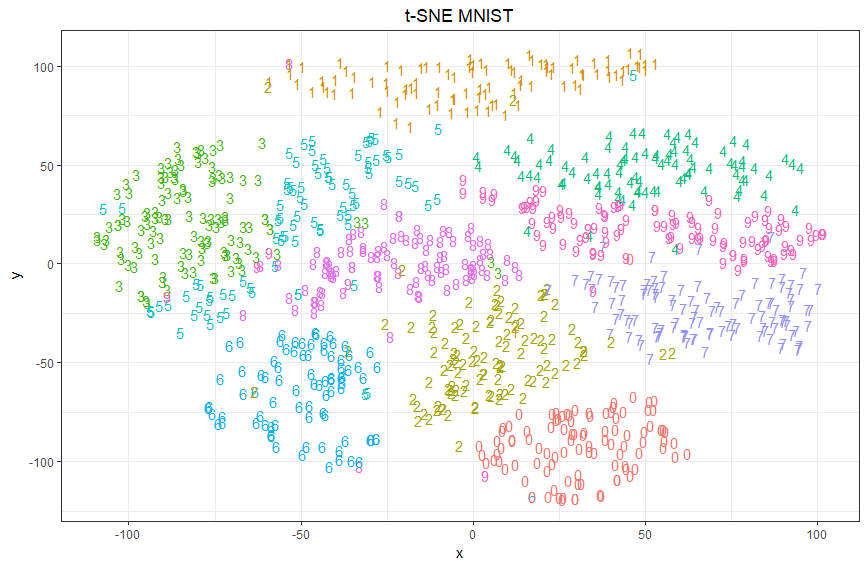
Pro: I was able to pack an extra dimension into each point by using a glyph to represent each point. It’s easy to see that predictions form contiguous regions in the 2D space.
Con: Misses can only be seen by carefully scanning the image for digits with the wrong color. Because only a sample of data is shown and there are relatively few misses, it is unclear exactly where the decision boundaries are.
2D Kernel Density Plot
To correct these defects, I next moved away from a scatter plot of the sample and looked for a way to visualize the underlying distribution. One way to do this is to compute a two-dimensional kernel density estimate from the underlying data and to use a contour plot to display the result. Essentially, we get a “hill” for the region where a particular class is prevalent. These regions look like concentric rings, although the rings are very far from circular. The probability that a given point belongs to any particular class is proportional to the number of rings of the right color that completely surround that point. Points which are in the intersection between two regions are ambiguous and this is where we should expect to see the most misclassifications.
centers <- xy %>%
group_by(label) %>%
summarise_all(funs(mean))
ggplot(xy[1:1000,], aes(x, y, color=label)) +
geom_density_2d() +
xlim(-120, 125) +
ylim(-130, 110) +
geom_text(
data=centers,
aes(x, y, color=label, label=label),
fontface="bold",
size=10,
show.legend=FALSE
) +
theme(legend.position="bottom") +
guides(colour = guide_legend(nrow = 1)) +
ggtitle("t-SNE MNIST, 2D Kernel Density")
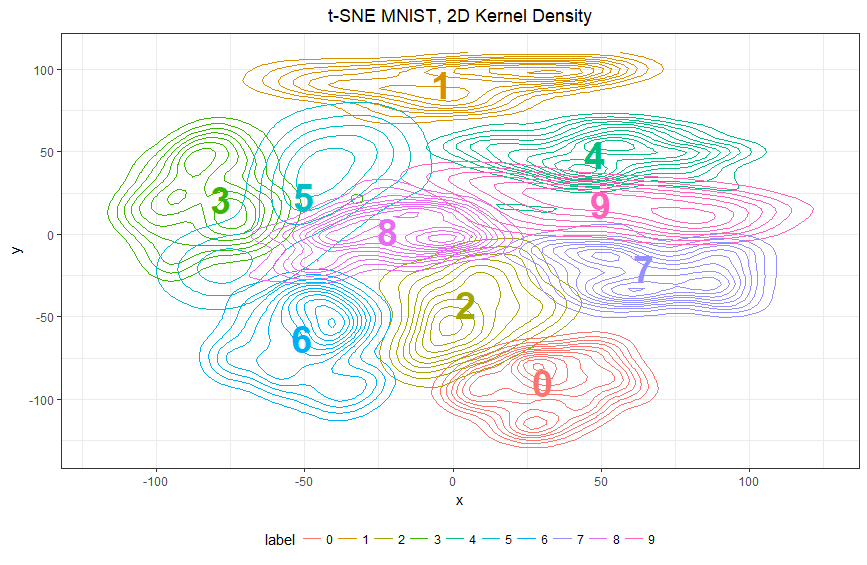
Pro: the simple expedient of labelling each centroid with a large color coded digit makes works well and makes the color legend at the bottom almost unnecessary.
The graph is highly interpretable at a glance, and can also be used to give precise predictions if you are patient enough to count rings.
It is very easy to see where boundaries overlap and the classification model may be confused: not the large area of overlap between 3 and 5 on the left, for example. Such overlaps directly correspond to pairs of classes for which misclassification is common.
Con: Directionality is rather unclear - if a point is in the overlap of 3 and 5, it is at risk of being misclassified - but will 3’s be misclassified as 5’s, vice versa, or both? Also, people do need at least some training to interpret contour plots, especially overlapping contour plots, which are not very common at all.
Hits and Misses Plot
Returning to the scatterplot concept and striking out in a different direction, my next idea was to draw attention to misses by color coding by accuracy instead of by class; in the below plot: correct predictions are labeled in blue, incorrect in red.
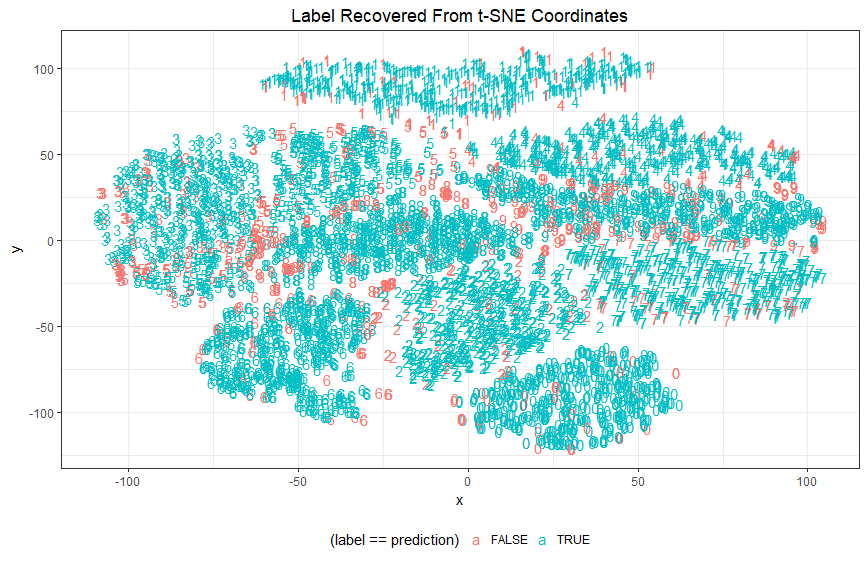
Pro: this variation naturally calls the eye to the misses: Unlike our first scatterplot the misses now stand out vividly.
It many ways this trick is successful: we can immediately see at a glance that misclassifications do indeed tend to fall near the boundary of two clusters, and we also get a sense of where such misses tend to belong to one class or the other. Finally, we can also easily pick out examples of misclassifications buried deep within other clusters - such cases are perhaps very far beyond the current model to correctly classify and represent the irreducible error of the current approach.
Con: Obviously we gave up the detailed information about the predicted class that we previously encoded into the color. Many of the criticisms directed at the previous scatterplot still apply here too.
Tree Plot
While some of the above visualizations have succeeded at attracting attention to the signal in the noise, they cannot be said to have algorithmically extracted the relevant information. The best way I came up with for doing this was to use hierarchical clustering which is sometimes used for similar problems, such as finding correlation relationships in a data set.
To apply the algorithm to this problem, I defined classes as “closer” to each other each other the more often they are misclassified as each other. If the algorithm does its job then those classes which are most likely to be mistaken together will be close together on the resulting tree. (Graphical plots of tree structures have the slightly pretentious name of “dendrograms,” terminology I will never-the-less adopt for precision.)
miss_table <- table(misses$label, misses$prediction)
sym_miss_table <- as.matrix(prop.table(miss_table + t(miss_table)))
diag(sym_miss_table) <- 0.07
sym_dist_table <- round(0.07 - sym_miss_table,4)
miss_dist <- as.dist(round(0.07 - sym_miss_table,4))
plot(hclust(miss_dist, method="ward.D"))

Pro: I am very pleased to note that this clustering is fundamentally successful: it correctly pairs 3 with 5, 4 with 9, and so on. These are the same patterns we observed in the less rigorous analysis above, but we no longer have to rely on eyeballing the graph and making a subjective judgement. The clustering algorithm is explicitly telling us that those are the most prevalent relationships.
Con: The above relationships are symmetric (as is required by the definition of a metric.) The use of a symmetric metric was in turn a requirement of the agglomerative clustering algorithm we used. We will need a fundamentally different approach for deal with directionality.
Heat Map
The dendrogram does obscure some of the raw data about the frequency of misclassifications, however. A standard way to have our cake and eat it to – to show both the algorithmic clusters and underling data in the same visualization – is to use a heatmap for the raw data, and attach the dendrograms to the rows and columns.
heatmap(sym_dist_table)
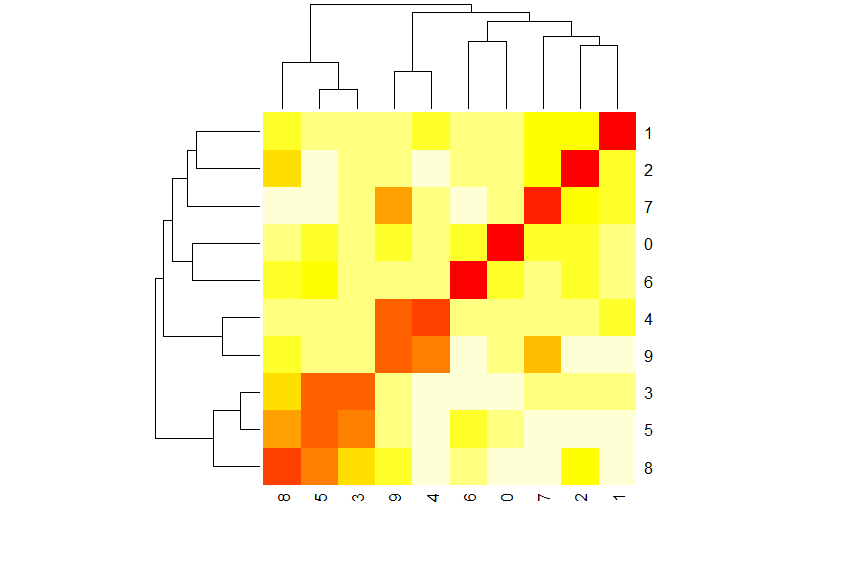
Pro: If you look closely, you’ll see that both the row and column dendrograms are in fact the same dendrogram from before, now being used to order the rows and columns of a heat map. This brings the 10 classes into a roughly block matrix form where squares along the diagonal indicate groups of classes that may be mistaken for one another. But the heatmap shows much more than this - we can see the isolated, bright red squares along the diagonal in the upper right, representing the easily classifiable cases. We can see not just pairs, but larger groups - the 3-5-8 group in the lower left stands out as a 3x3 block of related classes.
Con: I am very pleased with this visualization, and feel the only thing lacking is the directionality information we had to discard in order to fit our data into the hierarchical clustering mold. Let’s address that next.
Directed Graph
Let’s address the directionality issue now by returning to the asymmetric results matrix, before we applied the symmetry condition, and instead interpret it as the adjacency matrix of a directed graph. Then the classes will be the nodes of the graph and the edges will indicate common misclassifications. We can use the igraph package to visualize this digraph.
library(igraph)
plot(
graph_from_adjacency_matrix(t(miss_table > 12), mode='directed'),
main="Digraph: Real -> Mistaken Prediction")
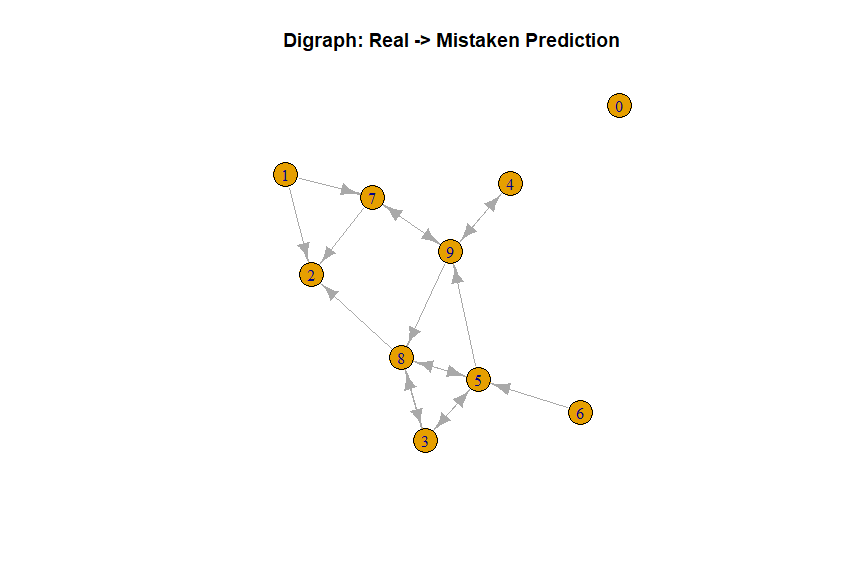
Pro: It makes certain imbalances in misclassification quite evident: while a 5 might be misclassified as a 9, a 9 will almost never be misclassified as a 5. Such imbalances can be found simply by looking for edges with only one arrow.
Con: Quite difficult to explain to a lay audience. No real sense of relative probability of each type of error. This graphic does not stand by itself, but may be a useful companion to the heatmap if directionality is present and relevant. While not necessarily a bad thing in and of itself, it does mean that we discarded directionality information.
Deep Dive into Misses
The above hierarchy suggests a strong relationship between the classes 3 and 5. We can explore this in depth by taking a random sample of such misses and plotting them in full.
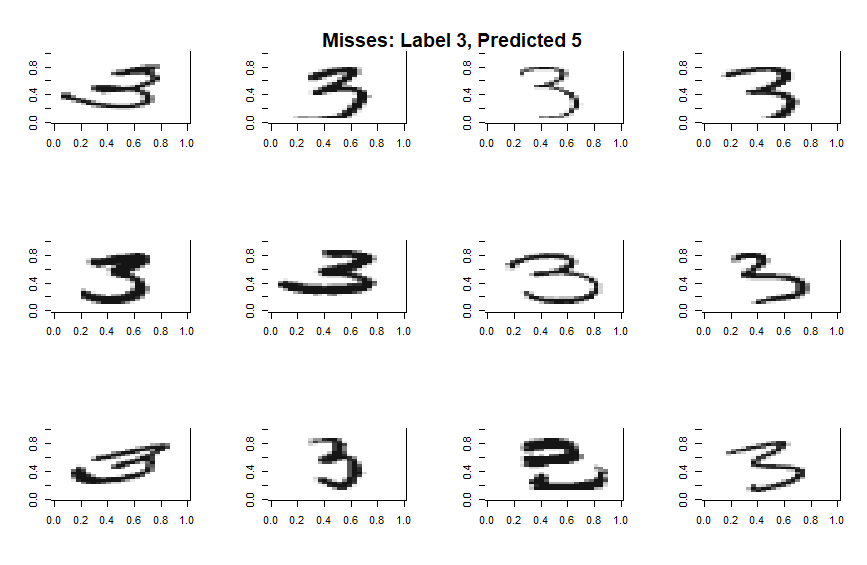

Some of these errors are more forgivable than others, but it’s clear that the multinomial algorithm is struggling when a digit is written in such a way as to shift critical features by a few pixels. An algorithm that didn’t look at all 784 pixels at once but zoomed in and looked for certain features or patterns in a translation invariant way would do a much better job… While I’m not too interested in the particulars of the toy problem, the fact that way to improve the model is immediately leaps to mind just by looking at a few examples of misses suggest that this kind of deep dive is a useful diagnostic supplement.
Conclusion
Performing hierarchical clustering on the $k \times k$ confusion matrix and displaying the results as a dendrogram was very successful at algorithmically finding real relationships between classes but hides directionality information. However, this can be supplemented with a digraph if directionality is important. I also found that presenting the dendrograms together with a heat map is an excellent way to visualize both the structure and raw results of a multiclass classification algorithm. Finally, I found that even a few concrete examples of each type of hit or miss went a long way towards providing insights about which cases the classifier could handle and which it could not.